Introduction
Bladder cancer is a heterogeneous disease with multiple molecular subtypes, each associated with distinct biological behaviors and clinical outcomes. Among these, Muscle-Invasive Bladder Cancer (MIBC) represents the most aggressive form, characterized by the tumor penetrating the muscular layer of the bladder wall. Approximately 10 to 20% of initially non-invasive bladder cancers eventually progress to MIBC, which is linked to a significantly higher mortality rate.
Recent studies have identified six molecular subtypes of bladder cancer — Luminal Papillary (LumP), Luminal Non-Specified (LumNS), Luminal Unstable (LumU), Stroma-Rich, Basal/Squamous (Ba/Sq), and Neuroendocrine-like (NE-like) — each requiring different therapeutic strategies and prognostic considerations. Accurate subtype classification is therefore crucial for personalized treatment planning.
In this project, we developed a Graph Attention Network (GAT) model to predict bladder cancer subtypes by integrating multi-omics data (such as gene expression profiles) and histopathological image features. By representing patients as nodes in a graph and leveraging attention mechanisms to capture relationships between molecular and visual features, our model learns complex biological interactions that contribute to more precise subtype classification.
This approach combines deep learning and graph-based reasoning to bridge the gap between image-driven and molecular-based cancer characterization, opening new perspectives for computational oncology and precision medicine.
Methods
In this project, we focused on classifying bladder cancer subtypes using a Graph Attention Network (GAT), and more specifically its improved version, GATv2. The graph was constructed based on the similarity between data samples, allowing the network to better capture relationships between patients through adaptive attention weights.
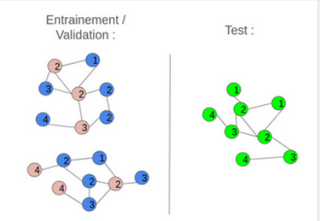
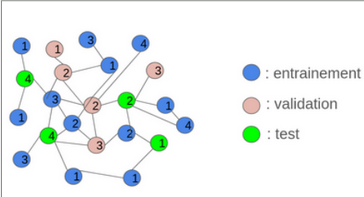
Two graph construction strategies were implemented: inductive and transductive learning. In the inductive approach, the initial graph was divided into several smaller, disjoint graphs for training and testing. This method offered faster computation and provided a good assessment of the model’s robustness on unseen nodes, although it resulted in some loss of global structural information. Conversely, the transductive approach used a single graph where some nodes were hidden during training. This preserved the complete graph structure and ensured high accuracy on data similar to the training set, but at the cost of higher computational complexity and lower generalization to new nodes.
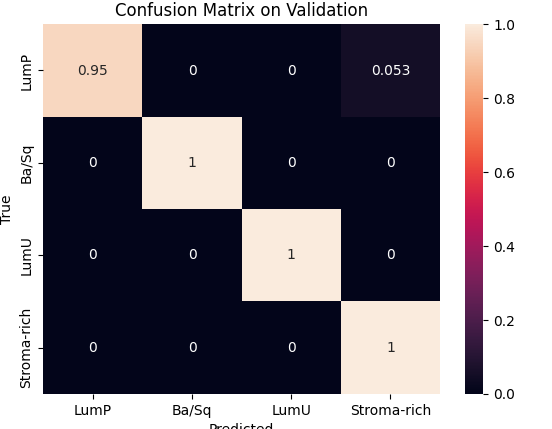
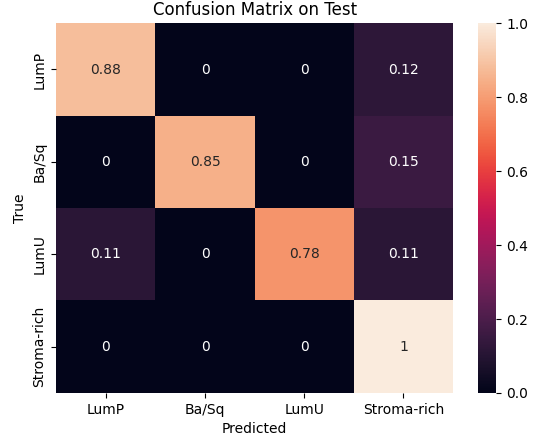
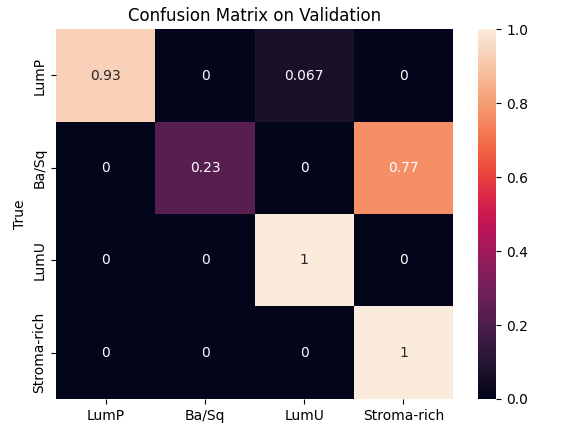
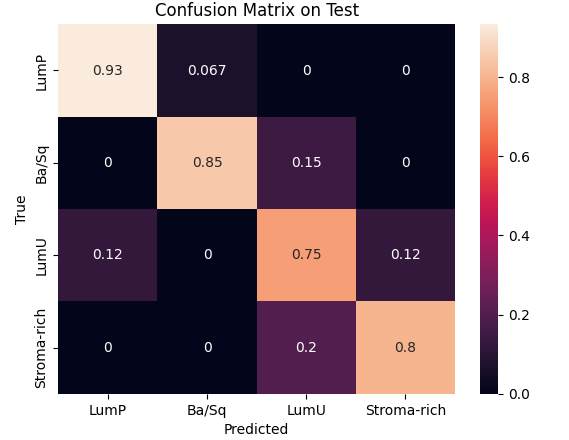
During preprocessing, we had to address a class imbalance problem among cancer subtypes. This required weighting each class to balance the model’s learning process and removing the two least represented classes to stabilize the results. We then used Optuna to optimize key hyperparameters such as the number of hidden channels and attention heads, with 10 and 25 trials respectively, each over 1500 epochs. The best configurations were 8 heads and 20 hidden channels for the inductive model, and 16 heads and 30 hidden channels for the transductive one.Finally, validation and testing sessions showed that the inductive model achieved the best trade-off between speed and performance, reaching 98 % accuracy on validation and 88 % on test, while the transductive model reached 80 % and 86 %, respectively.